P4
Characterization of the interaction of bacterial determinants and host cells during systemic inflammation for sepsis prediction
Summary
Sepsis may result in life-threatening organ dysfunction due to a dysregulated host response. However, no single biomarker has turned out to adequately reflect the rapidly evolving nature of a potentially septic patient’s status and to predict subsequent organ dysfunction. Hence, in this project, we want to focus on early physical changes and cellular effects, which are the sum of the complex host reactions rather than to look at singular markers to characterize bacteria-host interactions in the early stages of sepsis. Specifically, we plan to apply digital holographic microscopy (DHM), which is an interferometry-based stain-free technique that provides minimally-invasive, high resolution quantitative phase contrast imaging for live cell analysis, to characterize physical changes of lymphocytes and endothelial cells in the context of sepsis. DHM can detect fast morphological changes during time-laps investigations and captures physical cell parameters like volume, the refractive index and dry mass, indicating for example cell death processes. Furthermore, we want to use atomic force microscopy (AFM) to analyze nanomechanical changes of the endothelial glycocalyx (eGC), an up to 3 μm thick gel-like layer coating the luminal surface of the entire vascular endothelium. The eGC plays a pivotal role in the maintenance of microcirculatory homeostasis and is degraded in patients with systemic inflammation and sepsis.
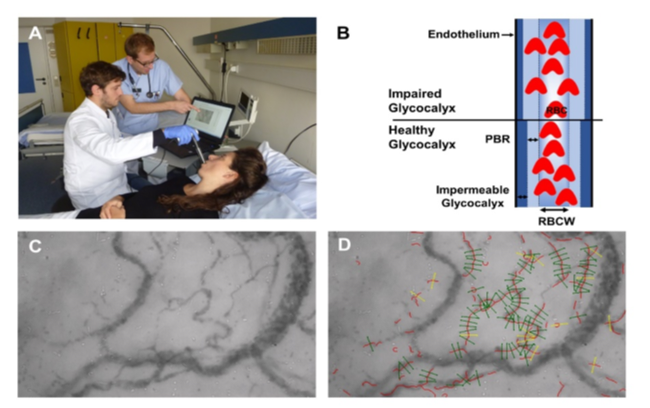
GlycoCheckTM System. (A) The GlycoCheckTM System consists of a sidestream-darkfield (SDF) camera coupled to a laptop. (B) Schematic illustration of cross section of a microvessel. GlycoCheckTM detects the dynamic lateral movement into the glycocalyx, which is expressed as the depth (in µm) of the perfused boundary region (PBR). (C) Representative image of the sublingual mucosa acquired with the SDF camera. (D) Quality check by the GlycoCheckTM software. Invalid vascular segments are marked yellow and are automatically discarded, while all valid vascular segments (green lines) are further analyzed. RBC: red blood cell, RBCW: red blood cell width.
In aim 1 we want to use DHM and AFM to characterize host cell reaction in vitro with focus on morphological/physical changes of lymphocytes and damage of the eGC. We hypothesize that host cells react depending on qualitative and quantitative differences among the bacterial determinants and that these reactions herald the onset and global net severity of a dysregulated host response.
In aim 2, we will translate our findings to the human system and investigate cellular changes of donor lymphocytes and endothelial cells exposed to retain blood samples from patients with microbiologically confirmed sepsis. Samples will be derived from the EDGE study (Early Detection of Glycocalyx Damage in Emergency Room Patients; Clinicaltrial.gov #: NCT03126032) within this consortium. Furthermore, we will prospectively measure sublingual eGC dimensions using intravital microscopy (GlycoCheckTM) and isolate lymphocytes for DHM analysis and in patients presenting to our ER with suspected infection. Our aim is to identify and validate specific DHM patterns and sublingual eGC values for early prediction of impending septic organ-failure (especially kidney and lung) and clinically relevant end-points. Finally, we will validate our findings in a second, prospective study, in which patients with a suspected infection requiring hospitalization are investigated for eGC-changes and morphological alterations in lymphocytes using GlycoCheckTM and DHM, respectively.





